Copper »
PDB 4tm7-4yst »
4unm »
Copper in PDB 4unm: Structure of Galactose Oxidase Homologue From Streptomyces Lividans
Protein crystallography data
The structure of Structure of Galactose Oxidase Homologue From Streptomyces Lividans, PDB code: 4unm
was solved by
A.K.Chaplin,
M.A.Hough,
J.A.R.Worrall,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 53.5 / 1.77 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 50.360, 126.630, 107.550, 90.00, 91.10, 90.00 |
| R / Rfree (%) | 18.767 / 22.819 |
Copper Binding Sites:
The binding sites of Copper atom in the Structure of Galactose Oxidase Homologue From Streptomyces Lividans
(pdb code 4unm). This binding sites where shown within
5.0 Angstroms radius around Copper atom.
In total 2 binding sites of Copper where determined in the Structure of Galactose Oxidase Homologue From Streptomyces Lividans, PDB code: 4unm:
Jump to Copper binding site number: 1; 2;
In total 2 binding sites of Copper where determined in the Structure of Galactose Oxidase Homologue From Streptomyces Lividans, PDB code: 4unm:
Jump to Copper binding site number: 1; 2;
Copper binding site 1 out of 2 in 4unm
Go back to
Copper binding site 1 out
of 2 in the Structure of Galactose Oxidase Homologue From Streptomyces Lividans
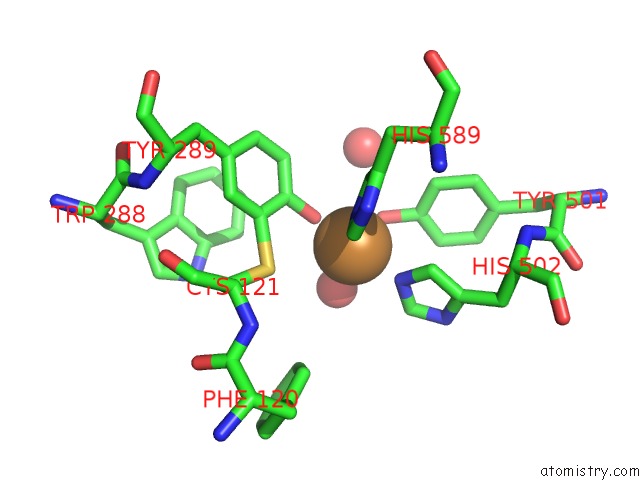
Mono view
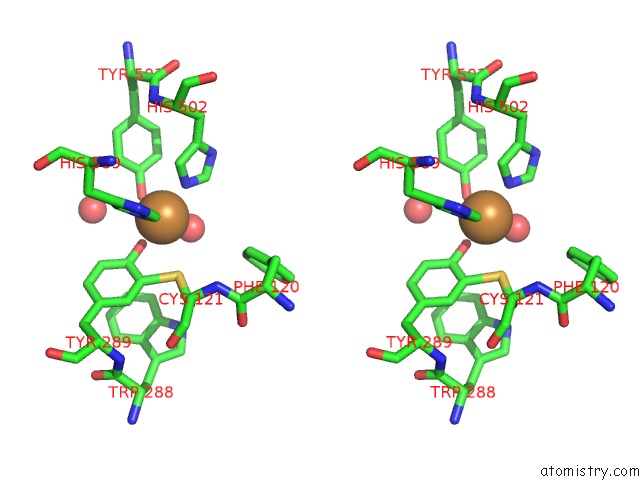
Stereo pair view

Mono view

Stereo pair view
A full contact list of Copper with other atoms in the Cu binding
site number 1 of Structure of Galactose Oxidase Homologue From Streptomyces Lividans within 5.0Å range:
|
Copper binding site 2 out of 2 in 4unm
Go back to
Copper binding site 2 out
of 2 in the Structure of Galactose Oxidase Homologue From Streptomyces Lividans
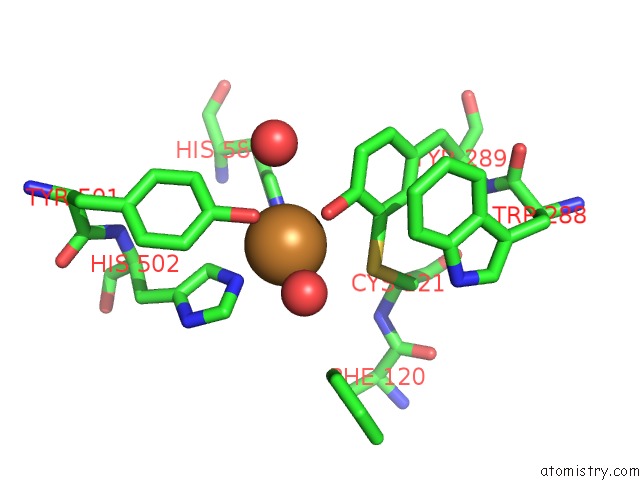
Mono view
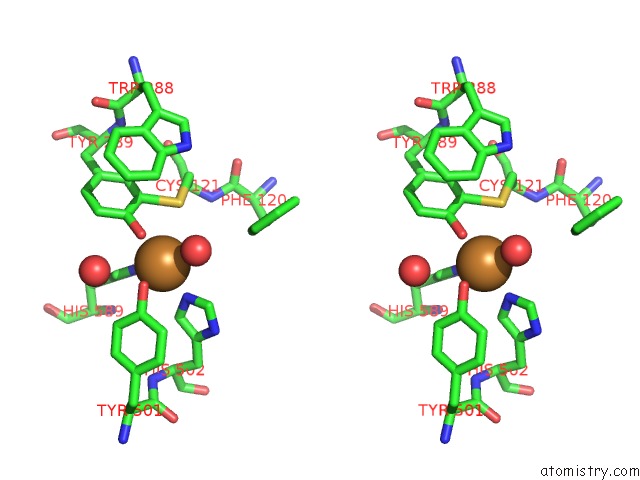
Stereo pair view

Mono view

Stereo pair view
A full contact list of Copper with other atoms in the Cu binding
site number 2 of Structure of Galactose Oxidase Homologue From Streptomyces Lividans within 5.0Å range:
|
Reference:
A.K.Chaplin,
M.L.Petrus,
G.Mangiameli,
M.A.Hough,
D.A.Svistunenko,
P.Nicholls,
D.Claessen,
E.Vijgenboom,
J.A.Worrall.
Glxa Is A New Structural Member of the Radical Copper Oxidase Family and Is Required For Glycan Deposition at Hyphal Tips and Morphogenesis of Streptomyces Lividans. Biochem.J. V. 469 433 2015.
ISSN: ISSN 0264-6021
PubMed: 26205496
DOI: 10.1042/BJ20150190
Page generated: Mon Jul 14 04:05:18 2025
ISSN: ISSN 0264-6021
PubMed: 26205496
DOI: 10.1042/BJ20150190
Last articles
F in 4IQVF in 4IQW
F in 4IQT
F in 4IQU
F in 4INB
F in 4IKT
F in 4IJU
F in 4IKL
F in 4IJV
F in 4IKK