Copper »
PDB 4hhw-4mai »
4lsy »
Copper in PDB 4lsy: Crystal Structure of Copper-Bound L66S Mutant Toxin From Helicobacter Pylori
Protein crystallography data
The structure of Crystal Structure of Copper-Bound L66S Mutant Toxin From Helicobacter Pylori, PDB code: 4lsy
was solved by
B.J.Lee,
H.Im,
C.C.Pathak,
H.J.Yoon,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 29.04 / 1.90 |
| Space group | P 1 21 1 |
| Cell size a, b, c (Å), α, β, γ (°) | 26.740, 83.998, 41.499, 90.00, 104.36, 90.00 |
| R / Rfree (%) | 19.9 / 26.1 |
Copper Binding Sites:
The binding sites of Copper atom in the Crystal Structure of Copper-Bound L66S Mutant Toxin From Helicobacter Pylori
(pdb code 4lsy). This binding sites where shown within
5.0 Angstroms radius around Copper atom.
In total only one binding site of Copper was determined in the Crystal Structure of Copper-Bound L66S Mutant Toxin From Helicobacter Pylori, PDB code: 4lsy:
In total only one binding site of Copper was determined in the Crystal Structure of Copper-Bound L66S Mutant Toxin From Helicobacter Pylori, PDB code: 4lsy:
Copper binding site 1 out of 1 in 4lsy
Go back to
Copper binding site 1 out
of 1 in the Crystal Structure of Copper-Bound L66S Mutant Toxin From Helicobacter Pylori
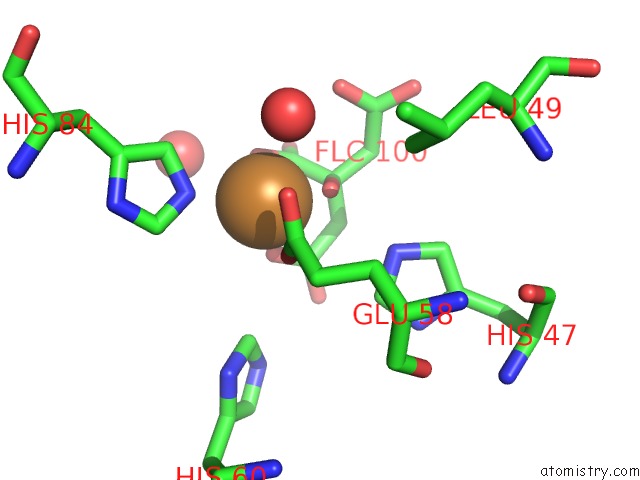
Mono view
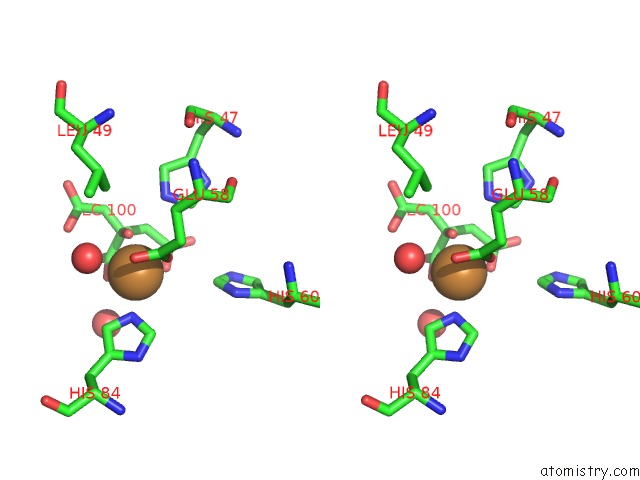
Stereo pair view

Mono view

Stereo pair view
A full contact list of Copper with other atoms in the Cu binding
site number 1 of Crystal Structure of Copper-Bound L66S Mutant Toxin From Helicobacter Pylori within 5.0Å range:
|
Reference:
C.Pathak,
H.Im,
Y.J.Yang,
H.J.Yoon,
H.M.Kim,
A.R.Kwon,
B.J.Lee.
Crystal Structure of Apo and Copper Bound HP0894 Toxin From Helicobacter Pylori 26695 and Insight Into Mrnase Activity Biochim.Biophys.Acta V.1834 2579 2013.
ISSN: ISSN 0006-3002
PubMed: 24060809
DOI: 10.1016/J.BBAPAP.2013.09.006
Page generated: Wed Jul 31 03:13:27 2024
ISSN: ISSN 0006-3002
PubMed: 24060809
DOI: 10.1016/J.BBAPAP.2013.09.006
Last articles
Cl in 3DOYCl in 3DP0
Cl in 3DOZ
Cl in 3DOL
Cl in 3DOK
Cl in 3DOJ
Cl in 3DNU
Cl in 3DNA
Cl in 3DN1
Cl in 3DN6