Copper »
PDB 2idq-2pp7 »
2k1r »
Copper in PDB 2k1r: The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I)
Enzymatic activity of The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I)
All present enzymatic activity of The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I):
3.6.3.4;
3.6.3.4;
Copper Binding Sites:
The binding sites of Copper atom in the The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I)
(pdb code 2k1r). This binding sites where shown within
5.0 Angstroms radius around Copper atom.
In total only one binding site of Copper was determined in the The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I), PDB code: 2k1r:
In total only one binding site of Copper was determined in the The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I), PDB code: 2k1r:
Copper binding site 1 out of 1 in 2k1r
Go back to
Copper binding site 1 out
of 1 in the The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I)
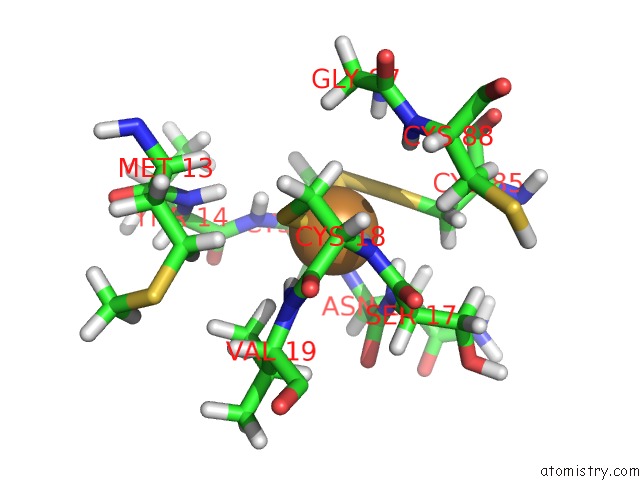
Mono view
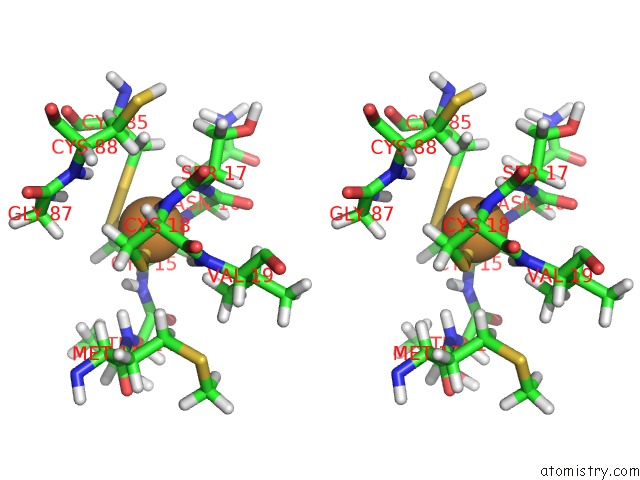
Stereo pair view

Mono view

Stereo pair view
A full contact list of Copper with other atoms in the Cu binding
site number 1 of The Solution uc(Nmr) Structure of the Complex Between MNK1 and HAH1 Mediated By Cu(I) within 5.0Å range:
|
Reference:
I.Bertini,
L.C.Banci,
I.C.Felli,
A.Pavelkova,
A.Rosato.
The Solution Structure of the Copper(I)-Mediated Complex Between the First Soluble Domain of the Menkes Protein and the Metallochaperone HAH1. To Be Published.
Page generated: Mon Jul 14 01:14:33 2025
Last articles
F in 4CCUF in 4CCB
F in 4CBT
F in 4CAV
F in 4CAP
F in 4CAR
F in 4CAN
F in 4CAO
F in 4C8B
F in 4C73