Copper »
PDB 1oe2-1rjp »
1phm »
Copper in PDB 1phm: Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat
Enzymatic activity of Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat
All present enzymatic activity of Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat:
1.14.17.3;
1.14.17.3;
Protein crystallography data
The structure of Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat, PDB code: 1phm
was solved by
S.T.Prigge,
L.M.Amzel,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 6.00 / 1.90 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 68.400, 68.660, 81.380, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.6 / 26.1 |
Copper Binding Sites:
The binding sites of Copper atom in the Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat
(pdb code 1phm). This binding sites where shown within
5.0 Angstroms radius around Copper atom.
In total 3 binding sites of Copper where determined in the Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat, PDB code: 1phm:
Jump to Copper binding site number: 1; 2; 3;
In total 3 binding sites of Copper where determined in the Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat, PDB code: 1phm:
Jump to Copper binding site number: 1; 2; 3;
Copper binding site 1 out of 3 in 1phm
Go back to
Copper binding site 1 out
of 3 in the Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat
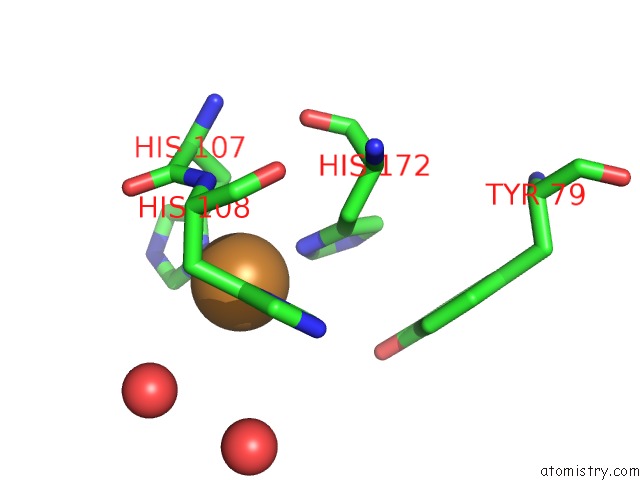
Mono view
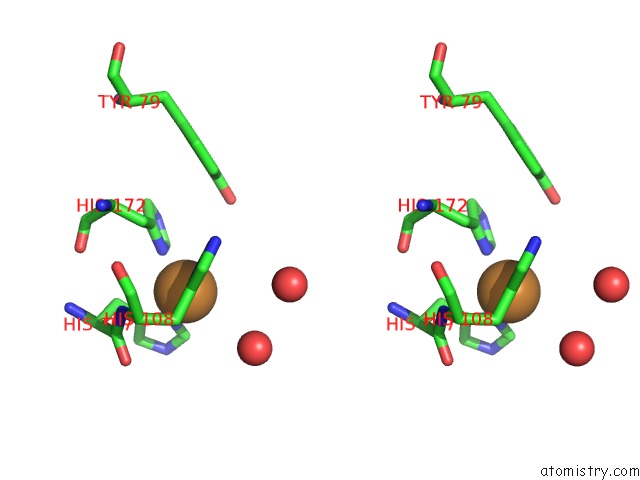
Stereo pair view

Mono view

Stereo pair view
A full contact list of Copper with other atoms in the Cu binding
site number 1 of Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat within 5.0Å range:
|
Copper binding site 2 out of 3 in 1phm
Go back to
Copper binding site 2 out
of 3 in the Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat
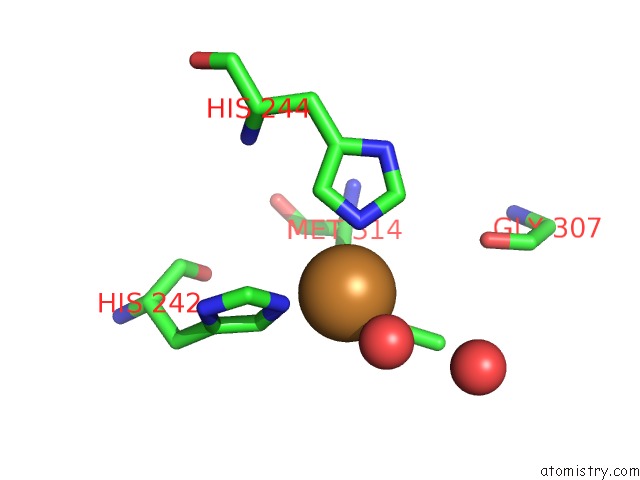
Mono view
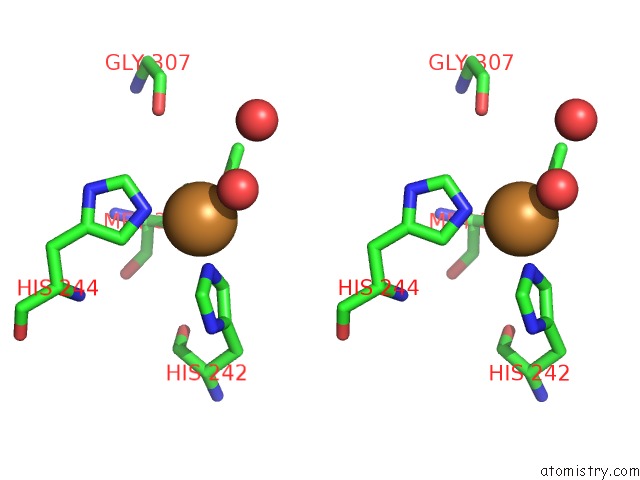
Stereo pair view

Mono view

Stereo pair view
A full contact list of Copper with other atoms in the Cu binding
site number 2 of Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat within 5.0Å range:
|
Copper binding site 3 out of 3 in 1phm
Go back to
Copper binding site 3 out
of 3 in the Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat
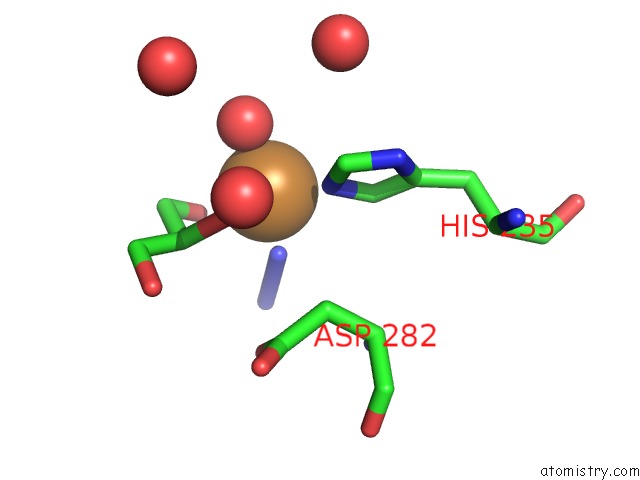
Mono view
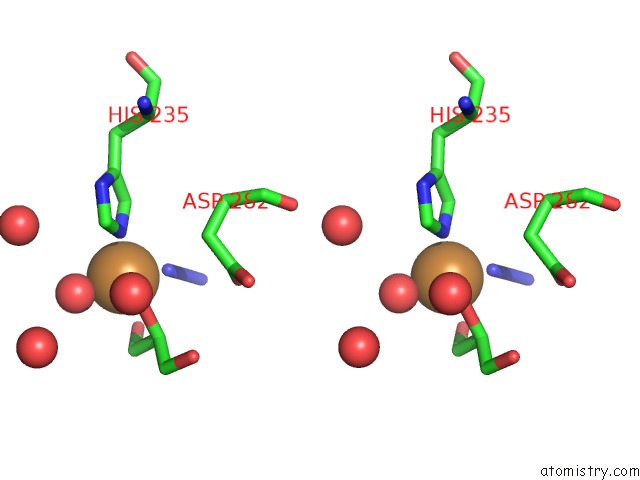
Stereo pair view

Mono view

Stereo pair view
A full contact list of Copper with other atoms in the Cu binding
site number 3 of Peptidylglycine Alpha-Hydroxylating Monooxygenase (Phm) From Rat within 5.0Å range:
|
Reference:
S.T.Prigge,
A.S.Kolhekar,
B.A.Eipper,
R.E.Mains,
L.M.Amzel.
Amidation of Bioactive Peptides: the Structure of Peptidylglycine Alpha-Hydroxylating Monooxygenase. Science V. 278 1300 1997.
ISSN: ISSN 0036-8075
PubMed: 9360928
DOI: 10.1126/SCIENCE.278.5341.1300
Page generated: Mon Jul 14 00:16:22 2025
ISSN: ISSN 0036-8075
PubMed: 9360928
DOI: 10.1126/SCIENCE.278.5341.1300
Last articles
F in 4IKFF in 4IJH
F in 4IGS
F in 4IF4
F in 4IIZ
F in 4IDQ
F in 4IGH
F in 4IGA
F in 4IBJ
F in 4IFY